|
The OMM exhibits consist of a two-framed web page. Molecules, figures, and text can be loaded into either of
the frames, depending on the instructional goals of the author. Embedded buttons in one frame can be used to invoke
RasMol/Chime scripts that affect a molecule being viewed by Chime in the same, or different, frame. The following
instructions describe a simple format that will allow you to build tutorial exhibits like those at the OMM. The
files described below, including a sample *.pdb protein, can be downloaded as a zip file, exhibit.zip.
Just save the file locally, unzip it, and modify the files as needed.
A) Build a directory for each molecular exhibit.
1. Use a file manager program (Explorer, say, for Windows 95/NT) to make a directory to contain the exhibit,
e.g. "exhibit". Into the exhibit directory place three other directories, named "frames", "images",
and "pdbs". The directory should look like this:

B) Download a *.pdb file from an appropriate source.
1. There are several ways of gaining access to molecular files for viewing by Chime:
- Construct an atomic coordinate file on your own using molecular modeling software, like Hyperchem. To
do this, you will need to purchase an appropriate software package.
- Visit Web sites that serve atomic coordinate files for internet access. You may either use your browser to
view the molecules, or download the molecules by using your right mouse button (Windows users) or the equivalent
Command key mouse button (Mac users) to activate the "Save As" option to save the file to a local directory
on your computer. See Links of interest.
- Visit Web sites that provide molecules for downloading. A comprehensive collection of macromolecules for which
atomic coordinates have been determined or modeled can be found at the Protein Data Bank (PDB). See Links
of interest.
2. Save the molecular coordinate file in text format, as a *.pdb file, e.g. "molecule.pdb". Save this
file into the "pdbs" directory (see above). This file will now be available for viewing by Chime in your
web browser.
C) Construct a "master" *.html (*.htm) file that builds the frames of the
exhibit.
1. Using either a text editor, or a web page editor, paste the entire following HTML code (<html> - </html>) into a file (note - comments will only
appear in the editor, not in the page as read by a web browser):
------------------------------------------------------------------------------------------------------------------------------------------------------
<html>
<head>
<title>Exhibit Name</title>
</head>
<!--Comment: the following code builds a two-framed web page
that will display a molecule in the left frame and tutorial text in the right frame. The molecule display will
occupy 60% of the width of the web page. The text frame will occupy 40% of the width of the page. -->
<frameset cols=60%,40%>
<!-- Comment: left frame--> <frame name="molecule" scrolling=auto src="frames/molecule_name.htm">
<!-- Comment: right frame--> <frame name="text" scrolling=auto src="frames/text.htm">
</frameset>
</html>
------------------------------------------------------------------------------------------------------------------------------------------------------
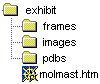
2. Save this file as a text file into your exhibit (root) directory and name it something
like "molmast.htm" (for molecule master). Your directory structure now looks like:
D) Construct a molecule *.html (*.htm) file for the left frame that loads the molecule
(*.pdb file).
1. Paste the following HTML code into a file:
------------------------------------------------------------------------------------------------------------------------------------------------------
<html>
<!--Comment: the background color of the frame will be black.-->
<body bgcolor="#000000">
<!--Comment: the following code retrieves the *.pdb file from the "pdbs"
directory. The molecule will spin 20 degrees/sec around the y axis upon loading.-->
<embed src="../pdbs/molecule.pdb" align=abscenter width=100%
height=97% spiny=20 startspin=true color3d=cpk display3d=wireframe name="molecule_name">
</body>
</html>
------------------------------------------------------------------------------------------------------------------------------------------------------
2. Save this file as a text file into your "frames" directory and name it "molecule_name.htm"
(this name must be the same as the *.htm file specified in the the left frame of your "molmast.htm" file
- see C, above).
E) Construct a text *.html (*.htm) file for the right frame that contains tutorial
text and buttons that run Chime scripts to change the rendering of the molecule displayed in the left frame.
1. Paste the following HTML code into a file (note- the embedded button invokes a sample
script - to access scripting tutorials and manuals see Links of interest):
------------------------------------------------------------------------------------------------------------------------------------------------------
<html>
<!--Comment: the background color of the frame will be black,
text will be white, and all links will be purple.-->
<BODY BGCOLOR="#000000" TEXT="#F9F9F9" LINK="#A217FD" VLINK="#A217FD"
ALINK="#A217FD">
<!--Comment: type in normal text as desired.-->
<!--Comment: the following code inserts a button into your page. The button
is 20 x 20 pixels and when clicked on will run the Chime commands indicated between the script " ". The commands change the molecule from a wireframe display
of all atoms to a ribbons display of the molecule backbone. Alpha helices will be colored magenta and beta strands
will be colored yellow. The molecule will be rotated 360 degrees around the x axis in 8 seconds. Note: the target
refers to the frame name of the left frame, as designated in the molmast.htm file-->
<embed type="application/x-spt" width=20 height=20
align=absmiddle button="push" target="molecule"
script="
select all;
wireframe off;
ribbons;
select helix;
color magenta;
select sheet;
color yellow;
move 360 0 0 0 0 0 0 0 8">
<!--Comment: type
in normal text as desired.-->
<!--Comment: the following code makes the indicated
hypertext load a different *.htm file (molecule_name2.htm)
into the left frame. This new *.htm file (in the frames directory)
might contain an embed command to load a different *.pdb file than
the original (molecule_name.htm - see D, above). A better way to
load new a *.pdb is shown in the Introduction
to Chime Scripting exhibit. -->
As can now be seen,
<a href="molecule_name2.htm" target="molecule">new molecule</a> contains significant structural homology to...
</body>
</html>
------------------------------------------------------------------------------------------------------------------------------------------------------
2. Save this file as a text file into your "frames" directory and name it "text.htm"
(this name must be the same as the *.htm file specified in the the right frame of your "molmast.htm"
file - see C, above).
F) You are on on your way! Your "molmast.htm" file should now be viewable
with your browser, provided the Chime plugin has been installed properly (see Required
software).
|
