I. IntroductionThe mechanistic details of DNA synthesis in higher organisms is not completely understood, in part because so many molecules are involved in the process. The bacteriophage T7 DNA replication complex is a good model system for the study of the mechanism of DNA synthesis because it consists of relatively few proteins. At left is the crystal structure of a T7 DNA polymerase complex and a short stretch of double stranded DNA with the primer and template strands indicated. The polymerase is caught in the process of adding a nucleotide to the 3' end of the primer DNA strand. II. Structural FeaturesThe crystal structure of the polymerase complex reveals two proteins essential to replication of the phage genome:
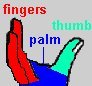
Like many other DNA polymerases, the T7 polymerase can be visualized as an open right hand, composed of a thumb domain that binds to thioredoxin, a fingers domain in which catalytic activity resides, and a palm domain.
The DNA is cradled in this open hand, with the palm domain forming a plate at the bottom of the cleft formed by the thumb and fingers domains. An N-terminal exonuclease domain abuts the palm domain. The polymerase domains are built mostly of alpha-helices, which play important roles in nucleotide recognition as well as in maintaining overall 3-D structure. Several domains also include beta-sheets. For example, the palm domain contains a prominent beta-sheet that forms the bottom of the DNA-binding cleft. There are several other key proteins involved in T7 viral DNA synthesis that are not shown in the structure at left: 1) a hexameric T7 primase-helicase that unwinds and primes the DNA; 2) a T7 single-stranded DNA binding protein that binds unwound, ssDNA in anticipation of DNA synthesis.
III. DNA SynthesisShown at left is a close up of the DNA primer strand (the daughter DNA strand that is being built by polymerization of nucleotides) and the template strand (the parental strand). Also shown is an incoming nucleotide (dGTP) that is being added to the growing primer strand. The last added nucleotide (dA) is indicated in yellow. Hydrogen bonding between complementary base pairs is indicated by dashed lines. Note the free 3'OH of the last added nucleotide on the primer strand (H not shown). During DNA primer extension, this hydroxyl oxygen attacks the alpha phosphorous atom of the incoming nucleotide in an SN2 reaction. This attack links the incoming nucleotide to the primer strand, resulting in the loss oftwo phosphates from the incoming nucleotide. A universal feature of nucleic acid polymerases is the use of two metal cations to catalyze the addition of new nucleotides to a growing chain. In the T7 DNA polymerase active site, two magnesium ions are positioned to facilitate the polymerization reaction. One Mg++ is juxtaposed to the 3' OH of the last added nucleotide on the primer strand. This stabilizes the ionized form of oxygen (O-), increasing its nucleophilicity, which leads to the SN2 attack on the alpha phosphorous atom. The other Mg++contributes to the reaction by stabilizing negative charges on the diphosphate leaving group. Now let's explore how these magnesium ions are positioned by T7 DNA polymerase. Three catalytic site residues of the palm domain (Asp475, Asp654, and Ala476) as well as several water molecules (H's not shown) associated with the finger domain coordinate the two Mg++ atoms in the heart of the DNA binding cleft of the polymerase.
IV. ReferencesDoublie S., Tabor S., Long A., Richardson C., and Ellenberger T.: Crystal Structure of a Bacteriophage T7 DNA Replication complex at 2.2 A Resolution. Nature 391: 251-258 (1998).
|
|